Machine Learning (ML), as a part of Artificial Intelligence (AI) methods, has considerably shaped the way we live in the past years. AI is incorporated in today’s search engines, marketing solutions, smart devices, social media, autonomous vehicles, drones, entertainment, industrial inspection, games, military… and as such, it has a profound impact on our lives. In all likelihood, this impact will only increase in the years to come. Slowly, but surely, AI is making its way to medical and clinical applications. Before we dive into specific AI applications and use cases, let us discuss for a moment what AI and ML are.
By dr. Danilo Babin, imec-TELIN-IPI, Ghent University, Belgium
What is AI?
The most simplistic way to think about AI is that it is a classifier (for readers with profound knowledge of ML: please take into account that the following explanation is aimed at non-experts and allow for some deviations from the scientific rigor on the topic). What does it mean? Simply, the classifier just assigns (classifies) the input data to some predetermined classes. For instance, a physician could use AI to examine images of suspicious skin lesions and classify them as “benign” or “malignant”. Another use case is to process images of the blood vessel in search of diseased vessels (e.g. aneurysms or malformations), where the vessels are eventually classified as “healthy” or “abnormal”.
How does ML work?
There are several ML principles and each of them has different characteristics. Some examples include Support Vector Machines, Decision Trees, Random Forests, and recently most popular Neural Networks (NN) such as Deep NNs, Convolutional NNs, Fully connected NNs…. The underlying concept of NNs resembles the functioning of a human brain in a very simplistic (but powerful) way. The whole principle is built on a “perceptron” – a unit (the main building block) of the NN. Perceptron is a basic element of a NN, which tries to mimic the behavior of a neuron in a human brain. The perceptron takes a number of input parameters and makes a decision based on the input values and weights assigned to each parameter (see Fig. 1). For example, let’s suppose I want to determine whether I should wear boots today. I base my decision on whether there is any rain (if it is raining the input value for this parameter is 1 otherwise the input value is 0), if it is cold outside (e.g. if it’s below 10 degrees I use input value 1 and if above the input value is 0) and whether I plan to be outside most of the day (input for “yes” is 1 and for “no” is 0). Now, if all 3 parameters are equally important, I assign them all a weight of 1/3 = 0.333… (all weights must sum up to 1) and for given circumstances (given input parameter values) perceptron computes the result by multiplying each input value with its weight, summing them all up and thresholding by value 0.5 (if above 0.5 it decides for wearing boots if below it decides against).
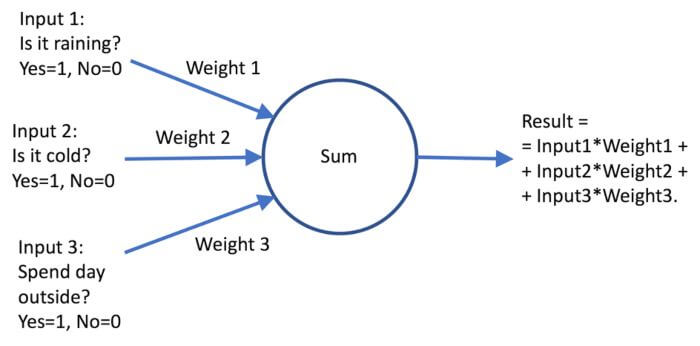
Fig.1: Perceptron
Although this is a simplistic example (no, I don’t need a perceptron to decide whether I will wear boots or not), it demonstrates how the perceptron works in a more complicated setting: the weights assigned to each of the inputs do not need to be the same. We can give priority to a certain input by increasing its weight and decreasing the weights of other inputs (e.g. if I would prioritize the “is it raining” parameter, I could increase its value to 0.4 and reduce the values of “is it cold” and “spending the day outside” parameters to 0.3. Note that if we increase the weight of the “is it raining” parameter above 0.5 the whole decision is based only on the fact if there is any rain or not). Indeed, in the above example, I set the weights of the perceptron myself, but in the case of a NN, the learning mechanism of the NN will adjust the weights of each perceptron individually according to the desired classification result on the training (sample) data. Note also that input values can be any values in a predefined range (not just 0 and 1, but any value in between), while the output is also a real value (again, not just 0 and 1, but any value in between, imagine it as a measure of the likelihood of a certain event). These generalizations make a perceptron a powerful decision system.
Creating a Neural Network (or “Building the brain”)
Up to now, we have explained the perceptron as the basic element of a NN, just as a neuron is for the brain (you can think of a perceptron as an equivalent of a “digital neuron”). The next step is to build a network using many perceptrons. Unlike the real human brain, which has many complex connections between its regions, we take a more simplistic way of constructing a NN. Let us imagine stacking many perceptrons into a layer (so, just vertically positioning many perceptrons one above the other) and connecting outputs of perceptrons from this layer to inputs of the next layer (so, we create another vertical stack of perceptrons behind the first layer and connect outputs of perceptrons from the first layer to inputs of the second layer perceptrons). If we continue adding layers of perceptrons we will end up with a network as depicted in Fig. 2.
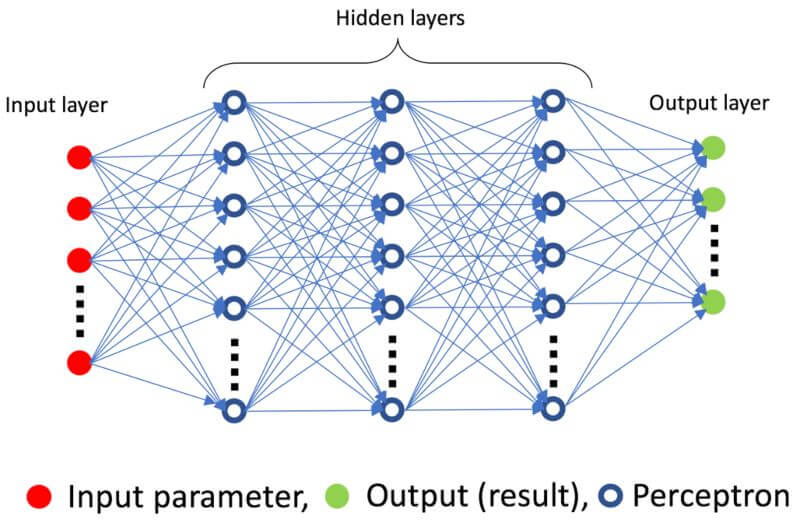
Fig.2: Neural network.
Each NN consists of:
- The input layer (input parameters from which we try to perform classification or in case of image processing each input node is a pixel from an image),
- Output layer (classes of classification, in case of skin lesion classification example, there is only 1 node in the output layer because we are working with a binary decision – the lesion is either “benign” and “malignant”, corresponding to results 0 and 1 on the output node)
- Hidden layer between input and output layers (hidden refers to the fact that their layers are not directly accessible to users). There can be multiple hidden layers of perceptrons (this is most often the case) and if we create a network of more than 1 hidden layer we talk of deep neural networks (there is some dispute about the number of layers needed to call a network a “deep” network, but let us say that 1 hidden layer of perceptrons will do) and if all perceptrons from one layer are connected to all perceptron in another layer we talk of a fully connected deep neural network.
What does this stacking of perceptrons in layers do for us? Each layer generalizes on the decisions made from perceptrons in the previous layer, which means that introducing more layers creates a network that should (if trained well) create more general rules for making classification decisions. The initial layers make “low level” decisions and as we move towards the end of the network, higher-level decisions are made. For example, let us consider how a NN could learn to recognize blood vessel branches and bifurcations in an image by starting from simple features and moving towards more complex (more general) features (see the illustration in Fig. 3).
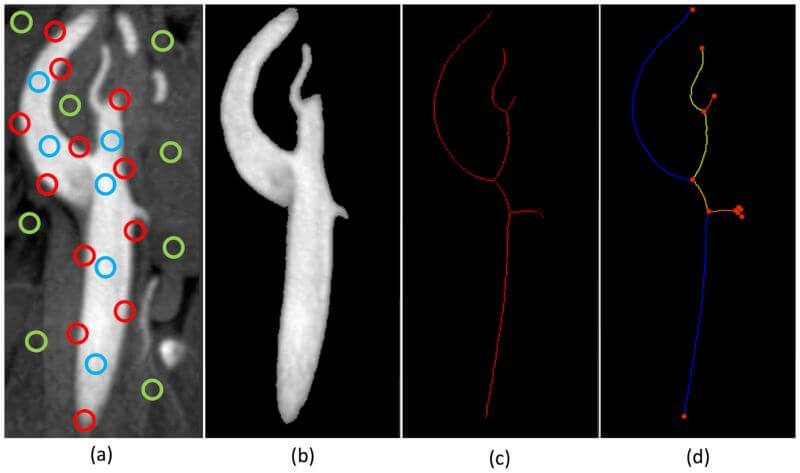
Fig. 3: Illustration of levels of learning in a NN: (a) extraction of simple features in the image (edges in red, regions of vessels in blue and background in green), (b) grouping of features into vessel segmentation, (c) extraction of centerline pixels from the segmentation and (d) division of the vessel tree into branches and bifurcations.
Part of this image is taken from DOI: https://doi.org/10.3174/ajnr.A1768
The first layer(s) of a NN would work on the low-level features, meaning that they would learn to extract edges (shown with red circles), regions of vessels (blue circles), and regions of background (green circles). The following layer(s) would learn to use the extracted features to extract the whole region of the vessel (i.e. perform segmentation). After the segmentation level, the following layer(s) would learn to generalize (simplify) the segmented structure by extracting centerline pixels. Finally, the last layer(s) would learn to interpret these centerlines to extract bifurcation points and vessel branches. Note how the network went from learning simple features of the image to interpreting and analyzing vessels as the final step. This is the type of generalization a NN does.
Another important thing to understand is that it is not enough to just create a powerful network, instead, the network has to be trained to perform a specific task. Training the network is just adjusting the weights of input parameters of perceptrons to fit a specific classification task. This is done by feeding the network with input data along with the desired classification result for each data sample. The network uses a learning method (Gradient Decent method) to adjust the weights of perceptrons automatically to obtain the best classification result on the training data. There is more to how the network performs learning, but this goes beyond the scope of this article (for readers interested in knowing more on the topic I would suggest looking up technical articles on neural networks [1,2]).
Data as a requirement for making a successful ML method
Despite their huge breakthrough in recent years, ML methods are not so novel and have been around for a few decades (they have, of course, improved significantly over that time). But, if ML is not so novel, the question which naturally arises is why it has made such a big breakthrough only now? There are multiple reasons for this, but the crucial one lies in the availability of training data. The key to designing an efficient ML method is having large quantities and wide variability of annotated data (annotated data means that for a specific data sample we exactly know to which class it belongs to, e.g. for an image of skin lesion we know with 100% certainty if the lesion is malignant or benign). This is called supervised learning because the ML model is learned from ground truth (annotated) examples. Unsupervised learning also exists (without annotated data), but extends out of the scope of this article. In either case, large quantities of data are required for the ML model to learn and to generalize. The reason for this is that the ML model needs to be presented with a representative sample subset for the specific class it is being trained. Simply, to encompass most of the variants within a specific class, we must use many training samples of that class. The most common (unanswered) question we encounter is “how much data do I need to efficiently train my ML method?”. The (unsatisfactory) answer is always “as much as possible”. A better question to think about is why is it so hard to determine the actual amount of data required for ML training? Multiple factors play a role here: type of the ML method (Deep Neural Networks, Convolutional Neural Networks, Random Forests, Support Vector Machines…), architecture (for NN: number of input, hidden and output layers, type of optimizers, batch size, number of epochs…), how many samples are required to make a representative subsample of the class of interest (e.g. if we try to segment a complicated shape it will need more samples than if we were segmenting something as simple as a sphere) and how different are the existing training samples (many training samples might be almost the same, so they do not hold any additional information).
Annotating the data
As we already pointed out, the availability of the data has allowed for a crucial step forward in the use of ML methods. Most of the hospitals already own huge databases of patient-specific data that are invaluable for training ML methods (of course, ethical approvals and patient consents must be acquired for such studies along with respecting GDPR). Very often the cumbersome process lies in annotating the training data. Specifically, this means that for each training sample we give a definitive label to which class it belongs. For example, in the case of skin cancer detection, we need to label each lesion image as being either “malignant” or “benign”. The image is being “fed” to the neural network together with the actual label (decision). In the case of skin cancer example, the label is determined from biopsy (excision of the lesion) and is 100% correct (the problem in AI-based skin cancer detection does not lie in the annotation, but in the fact that it requires tens of thousands of images to train the method). However, there are cases when the annotation is not perfect.
Image segmentation (a division of the image into objects of interest) is a process in which pixels of an image are classified as belonging to the object of interest (foreground) or everything else (background). For example, the segmentation of blood vessels will mark each pixel of blood vessels as foreground and everything else as background. This clearly equals the classification of image pixels into the foreground and background pixels, so using ML (as a classifier) is a valid way to go. The only problem is what to use as a label (ground truth). Ideally, we would want to have training data that is accompanied by an accurate ground truth – a map of which pixel belongs to which structure in the image. Such a (100% accurate) map does not exist simply because medical images are made on patients or volunteers. Checking if a pixel belongs to a certain organ or tissue of interest would require physically checking the structure and the anatomy inside the patient. Two possible solutions are given to resolve this problem. The first solution is the design of phantoms – artificial structures that resemble the organs or tissue of interest. In this case, we can know which pixel in the image belongs to which structure with 100% accuracy, but the problem is that the imaged structure is artificial. This means that the structure does not completely accurately resemble the actual (live) tissues or organs of interest. It is often very hard to accurately simulate the function of organs (blood flow, heartbeat,…). The second approach to annotating segmentation data is to use manual (handmade) annotations by experts. This involves multiple experts outlining the structure of interest in original images.
The process of manual annotations is flawed for 2 reasons: undetermined (questionable) accuracy and inter-observer variability between experts. The inter-observer variability relates to the fact that multiple experts will create different annotations (draw different regions for the same tissue/organ) on the same image. At the first glance, this might seem to be the case because of the difference in the level of experience, background, or seniority between the experts, but we now know that these do not play a crucial role. The same (one) expert annotating the same (single) image within a timespan of few months (or even weeks or days) between the annotations will annotate the image differently. Here we talk about intra-observer variability (variability of repeated annotations for a single expert). It is crucial to understand why this happens – why does the same expert annotate the same image differently in successive trials? Answering this question reveals also an underlying concept of ML methods. The answer is: because of learning.
Consequences of learning (or “should AI be used everywhere?”)
Learning is a concept of constantly updating our views and re-examining our principles for a specific matter at hand daily. Every morning we wake up with a slightly different view of our everyday problems and activities. Learning happens even when our mind seems to be resting – it uses the time off to gather our thoughts and create new concepts. With this in mind, we realize that the same person who annotates the same image multiple times does not in fact use the same principles in the annotation. Without even knowing, the person has changed (at least slightly) his or her views on the matter and created new concepts between the consecutive annotations, which result in the differences in annotations. The longer the period between annotations, the more likely is the annotation differences (and possibly more prominent). What does this have to do with ML? Does the same principle apply to ML methods? In fact – yes, it does.
Suppose that a neural network has been thought to resolve a certain classification problem on a set of training data, e.g. a neural network has been trained to segment malformations of cerebral blood vessels by using 100 annotated images and the trained method performs segmentation with decent accuracy. Now let us suppose that a year later we have acquired and annotated 100 more images. It makes perfect sense to re-train our method on the 200 images that we now have available to further improve the segmentation method. If we run our newly re-trained method on the same test data we used when we had only 100 images (this test data set is independent of the training data), we will notice that there are differences when comparing the current and earlier segmentation results. This is what we expected (because we introduced more images to improve the method), but the results may even be different for those cases where the segmentation was accurate in the earlier version (even worse, the results of the new method might even be worse than before, which indicates that overfitting happened – the method learned the training images by heart). Obviously, by re-training our method, we have hampered reproducibility, which is exactly what happens to human experts in their learning process. Even if we re-trained our network using the same set of data, the method would be trained differently. The reason is inherent for neural networks – the initial state of a perceptron is random and the learning process is also randomized (to allow for better generalization).
Does this mean that the accuracy of ML methods is at stake? Not at all. If trained well, the neural networks achieve high accuracy and high enough reproducibility. However, much depends on the annotations used, and we have shown that human annotations can be flawed. Does this mean that human expert annotations should not be used? No – human annotations can be of vital importance in cases where there is (almost) full agreement between the experts. However, for fine decisions (e.g. segmenting fine structures in images) and cases where reproducibility is of utmost importance, the classical (deterministic) image processing methods are still of high importance and should be preferred over methods trained on manual annotations (as such methods incorporate the mistakes contained in the flawed annotations). Here we come to a vital conclusion: a NN is only as good as the annotations used for its training and annotations based on human opinions can be flawed. To quote Agent Smith (“The Matrix”, 1999): “Never send a human to do a machine′s job”.
Use cases
We have already mentioned the use of AI in skin lesion assessment. In practice, the problem dermatologists encounter when assessing skin lesions is that they often send all suspicious lesions for biopsy (to make sure they don’t miss any malignant lesions), which results in many benign lesions being excised. Therefore, the goal of the AI can be defined as: ”lower the number of lesions marked for excision, without missing out on any malignant lesions”. In other words, we need to improve on specificity, while maintaining (or, if possible, improving) sensitivity.
The use case of blood vessel anomaly detection is a bit different in this aspect because the main goal here is to find all possible suspicious vessel regions and report them for a check-up by an expert. In this specific case, the expert has to take time to check all the vessels, which is a tedious and cumbersome process. Therefore, the goal of the AI is to point out the likely candidates for this examination without missing any real diseased vessels. In other words, the number of candidates that are presented to the expert for further examination needs to be high enough to include all diseased vessels. The fact that there are somewhat more candidates generated than required is not an issue, since the utmost goal is to detect the disease. Therefore, in this case, the AI aimed to increase the sensitivity at the cost of specificity.
Another interesting and often encountered use case in image processing is image segmentation. The segmentation is the delineation of a certain object (tissue, organ…) in the image for subsequent measurements. For example, if we want to measure the volume of a left ventricle of a heart, the common approach is to segment (delineate) the ventricle (determine which pixels in the image belong to the ventricle) and perform the measurement (count how many pixels belong to the ventricle, multiplied by the size of the pixel given in millimeters). In this context, the AI classifies the pixels in the image as “left ventricle” pixels or “other tissues” pixels. Most commonly used NNs include U-net architectures which have shown to be a great asset when the training data is scarce (and the usual data augmentation approaches do not manage to resolve the problem).
Detection of certain diseases is one of the most valued use cases for AI methods in medical practice. This is driven by constantly growing imaging capabilities and the availability of imaging devices. Full body MR imaging is starting to dominate over anatomy-specific MR imaging and the reason is that the full-body imaging still provides high enough detail (although imaging only a specific part of the body will, of course, result in more details), while having the advantage in capturing the overall state of the patient. Many related or even unrelated problems have been detected in patients using full-body MR. One problem that arises here is that such a large image needs to checked thoroughly and in high detail by a specialist (or even by multiple specialists) to detect the problems unrelated to the actual main health problem of the patient (which was the reason for MR examination in the first place). There is the need to have an automatic system that would check the full-body image for specific diseases and mark the suspicious areas for the follow-up by a specialist. Having a highly sensitive method is the goal in this case, as it would allow for most of the abnormalities to be detected, while it would still significantly reduce the examination time of the specialist.
The list of AI applications in medical practice is ever-growing, and with the introduction of smart wearable devices, AI is being used to predict certain health-related events from the wearable data (ECG, EEG, behavioral data…). With this approach, the goal is to enhance the prevention, which has shown to be critical in taking the load off the hospitals (better prevention means fewer treatment requirements) and improving the health of the general population.
Interpretability (or “is it all so great?”)
The main problem associated mostly with NNs is the interpretability of its results. Very often the results of AI confirm the observations of experts (especially if the training was performed on manually annotated data). However, in some cases, the opinions of AI and human experts differ. Normally, this isn’t anything unusual – opinions of multiple experts can differ on a specific topic, which is why multiple opinions are sought and the final opinion is given by consensus. However, in this process, it is of utmost importance for each expert to justify his or her opinion for that opinion to be taken into account. And this is where the NNs still fail to meet the requirements. NNs are still considered as a “black box” mechanism, and a lot of effort has been invested into making them interpretable (“white boxing”). To be clear, it is not true that we have no clue as to why certain classification results have been made – we can backtrack results to the inputs that had the most influence on the result. In the case of images, we can also determine which part of the image played a crucial role in making a certain decision. However, this is still not enough because of the fundamental difference in applied logic of human experts and the logic of the AI. For example, I have been recently involved in a Ph.D. topic aimed at localizing slices of functional ultrasound data within the anatomical structure of a rat brain. Even though the method performs well, the backtracking to prominent image regions shows that the most important part of the image in the case of AI was the region of small blood vessels, while all experts relied on exactly the opposite regions (outer regions where the shape of the brain and large blood vessels were visible) to perform the localization. The question here is – which is a better approach and what can we learn from the fact that the region of smaller vessels is used for classification by AI (besides the fact that that region is also a viable option)? It is hard to answer these questions because it seems that ML methods perform on a completely different level of logic compared to humans. There is still no clear way on how to map the logic of ML methods to the logic of humans. Until we resolve this problem, we will not be able to learn from the AI and we will not be able to fully trust it in situations of (critical) conflicting opinions.
The Future
The question that is often posed is whether the AI will replace physicians (and, eventually, all of us) in the future. Although it is impossible to predict the future, one thing has to be made clear – the goal of current AI developments in medical practice is to aid physicians in their everyday work (and not to replace them). Although AI has come a long way and is proving to be a powerful asset in many applications, it is by no means at the level to rival the capabilities of the human brain. AI has been built for solving classification problems and performs this profoundly well (much better or at least as good as humans), but requires a human expert to interpret and approve (or disprove) the results. The (fabulous) results achieved by the AI are often misinterpreted as being the product of an “all-powerful” mechanism that will resolve all our problems. This is, of course, an overstatement – AI, as it exists today, does not incorporate a profound understanding of the problem it is solving (this is manifested in cases where small changes in images have a high impact on the classification results, interested readers should look up “adversarial attacks on neural networks” [3]). AI is (still) not capable of translating its logic to human reasoning (which is a more robust one, as it incorporates multiple levels of prior information and intelligence). Instead, we should look at the AI as what it is – a very powerful tool that, if used well, will allow us to improve every aspect of our health system, from prevention to diagnosis and treatment.
